As the glow of festival lanterns fades out the cold old year and spring breezes bring joy across the land, Zhejiang University ushers in its first publication in Nature in the Lunar Year of the Horse, right in the joyous hum of New Year greetings.
Fatefully, the submission of this groundbreaking paper took place exactly one year prior, also on Lunar New Year's Day. After a year of unwavering dedication and groundbreaking breakthroughs, an interdisciplinary research team has birthed an entirely new technical approach: the G Protein-Coupled Receptor Exoframe Modulator (GEM).
What is GPCR, and why does it matter?
Known as the human body's largest "signal receiver", G protein-coupled receptors (GPCRs) are distributed across major tissues throughout the body. Through their seven-transmembrane helix structure, they regulate nearly all critical life processes, including sensation, emotion, cardiovascular function, and metabolism.
Targeting this pivotal signal hub, the research team led by Yan Zhang from the School of Medicine and Liangzhu Laboratory, and Min Zhang from the College of Computer Science at Zhejiang University, launched a cross-disciplinary collaboration and proposed an "exoskeleton-style" receptor regulation strategy for precise modulation of GPCR function. They fused the GEM module — dubbed the "exoskeleton armor" — with the native GPCR structure, expanding the classic seven-transmembrane helix into nine or even thirteen helices, thereby reshaping the receptor's signal output modes and corresponding biological functions.
Titled De novo Design of GPCR Exoframe Modulators, this achievement demonstrates artificial intelligence's practical capability in membrane protein design and functional reconstruction, offering a novel technical pathway for understanding and harnessing the regulatory potential of GPCRs.
The first author of the study is Shizhuo Cheng, a PhD candidate in the "Medicine +X" interdisciplinary program at the School of Medicine, Zhejiang University. Co-first authors include Jia Guo, Yunli Zhou, Xumei Luo from the School of Medicine, Zhejiang University, and Gufang Zhang from the School of Pharmaceutical Sciences, Soochow University. The corresponding authors are Yan Zhang and Min Zhang.
Screenshot of the paper on the Nature website

A Paradigm Shift in GPCR Research
As a core component of the cellular signal transduction system, GPCRs act as the cell's "signal relay stations" and are the therapeutic targets for more than 30% of globally approved drugs. During signal transduction, the unique seven-transmembrane helix structure of GPCRs undergoes a seesaw-like conformational change, converting external stimuli into intracellular biological signals.
"Currently, the vast majority of drug design targeting GPCRs centers on the orthosteric pocket—a natural ligand-binding site on these proteins," explains Yan Zhang. "This site can be seen as the receptor's 'on/off switch', with drugs exerting their effects by pressing or releasing this switch."
Clinically, however, hundreds of diseases caused by gene mutations lead to jamming or misalignment of the "components" of GPCRs, the body's vital signal receivers. For patients, such structural dysfunctions often result in long-term and intractable symptoms. Regrettably, traditional drugs designed around the orthosteric pocket struggle to repair these structural functional damages, frequently leading to limited therapeutic efficacy.
"There are more than 800 distinct GPCRs in the human body, yet only three to four transmembrane proteins have been confirmed to directly regulate GPCR function," notes Yan Zhang. This stark contrast led the team to realize that the extensive transmembrane regions of GPCRs still contain vast, unexplored territories that have not been systematically investigated.
This insight spurred a bold idea: instead of fixating on the orthosteric pocket—the traditional "switch"—the team would target the GPCR transmembrane domain, attempt to design artificial transmembrane proteins, and equip key receptor structures with a customizable, programmable "armor" to precisely modulate GPCR function.
"Since AI-generated text, images, and soluble proteins are now commonplace, why not use AI to generate transmembrane proteins too?" Yan Zhang recalled of the initial inspiration. It was against this backdrop that he partnered with Min Zhang, an AI expert from the School of Computer Science and Technology, to embark on an interdisciplinary exploration into the "uncharted territory" of membrane proteins.
AI de novo design workflow of GEM
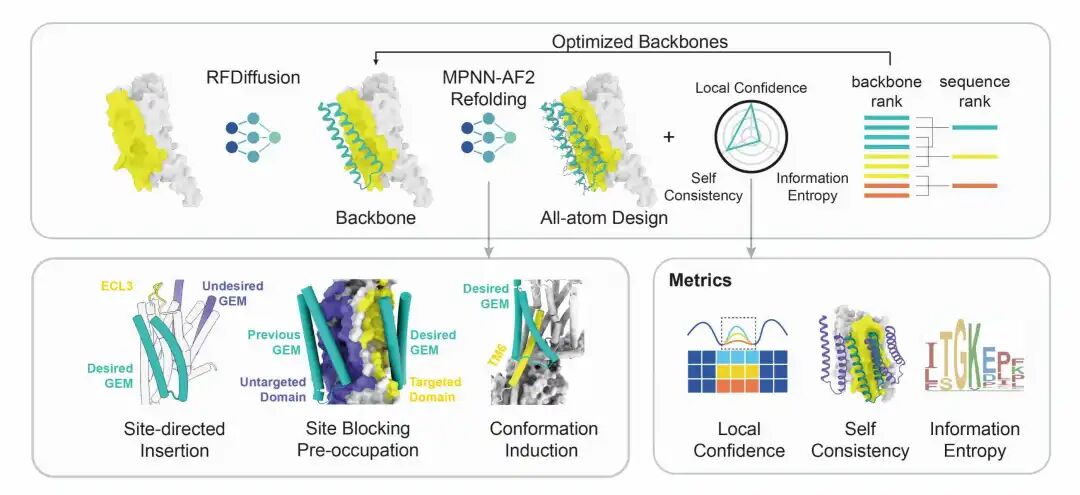
Innovative AI-Driven Design
De novo protein design—creating proteins with specific functions that do not exist in nature, without reference to native structural or sequence templates—is notoriously challenging in the field of protein engineering. Unlike modifying or upgrading existing proteins, this task puts researchers' and algorithmic models' understanding of proteins to the ultimate test.
"From a computer science perspective, we need to systematically decompose a seemingly simple design requirement into functional modules that AI can implement step by step," Min Zhang elaborates. Designing a protein is like building with blocks: constructing each layer incrementally to achieve the overall function.
Prior to this study, there were almost no mature examples of protein design targeting the transmembrane region of membrane proteins, leaving the team with no direct experience to draw on. The real challenge was not merely "creating a new protein", but first answering a more fundamental question: during GPCR's "sensation-conformational change-signal output" process, which transmembrane interfaces are actually involved in this mechanical movement, and which positions are most likely to influence signal direction? Only by understanding these structural mechanisms could the team determine where and how the "exoskeleton" should be attached.
Thus, GEM is no product of arbitrary "generation". The design originated from the team's years of accumulated research experience in GPCR structure and function. First, the team developed a computational "probe" to systematically scan the regions on the receptor's transmembrane surface that are inaccessible to traditional small-molecule drugs. This transformed the once obscure "uncharted territory" into an analyzable structural space, gradually identifying potential binding interfaces and regulatory nodes.
Point cloud map of dominant binding regions generated by the probe algorithm
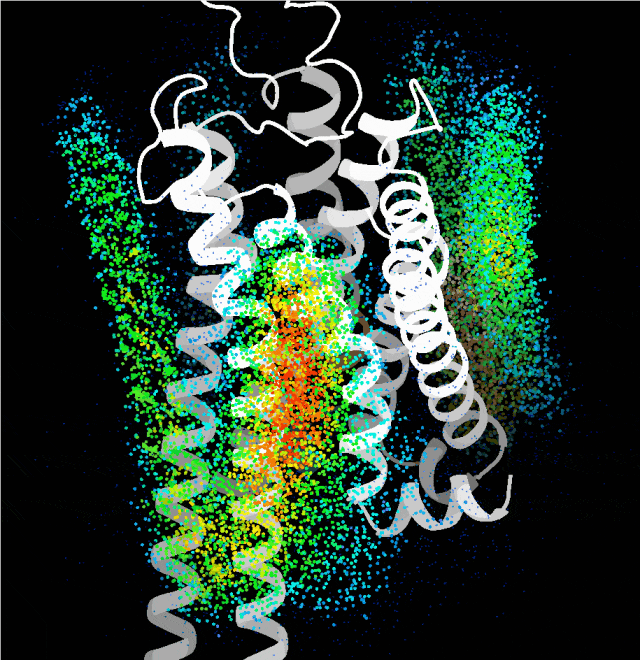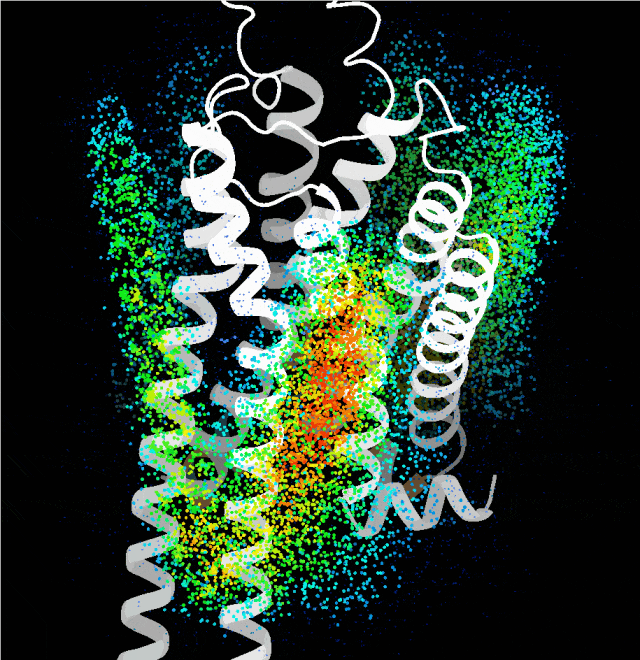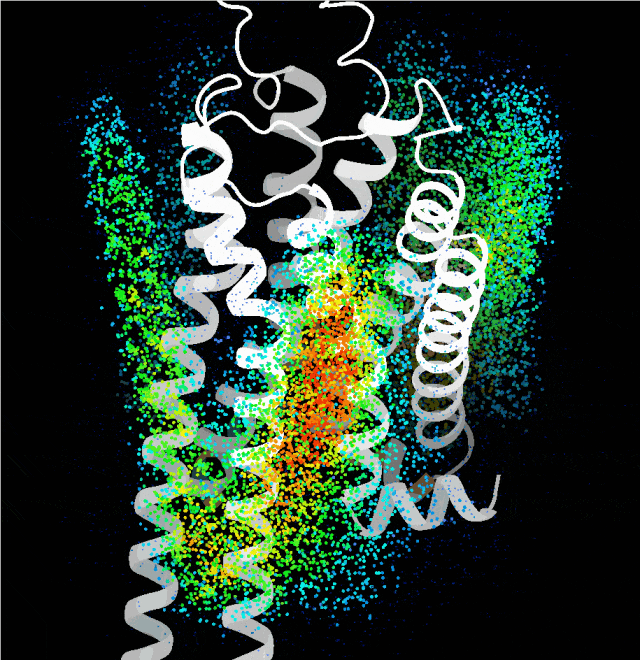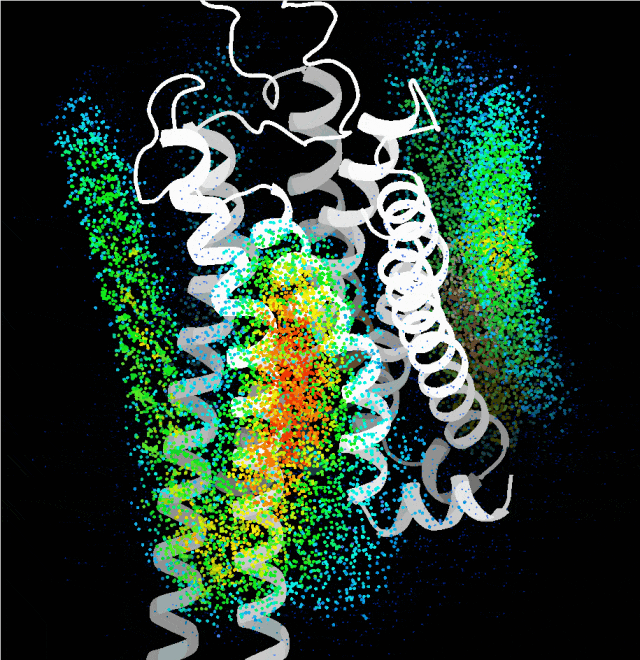
Building on this, the team integrated multiple complex factors including protein sequence, folding, binding sites, and conformation, and leveraged technologies such as generative diffusion models. They also taught the AI a strategy called "structural prompting", enabling it to generate high-confidence protein structures with distinct functions without any reference to native templates. The functional and structural rationality of these designs was further verified through cellular functional assays and cryo-electron microscopy structural analysis.
The two teams collaborated closely and advanced the research in tandem. Drawing on their solid expertise in GPCRs, the researchers developed three structural prompting strategies—site-directed insertion, pre-occupation site blocking, and functional conformation induction—centered on the key conformational changes in receptor signal transduction. Ultimately, they constructed transmembrane protein components that can precisely bind to specific epitopes of GPCRs and stabilize specific conformations, achieving diverse functions in experiments such as positive activation, negative inhibition, and biased regulation.
Functional experimental verification and structural analysis of different GEM variants
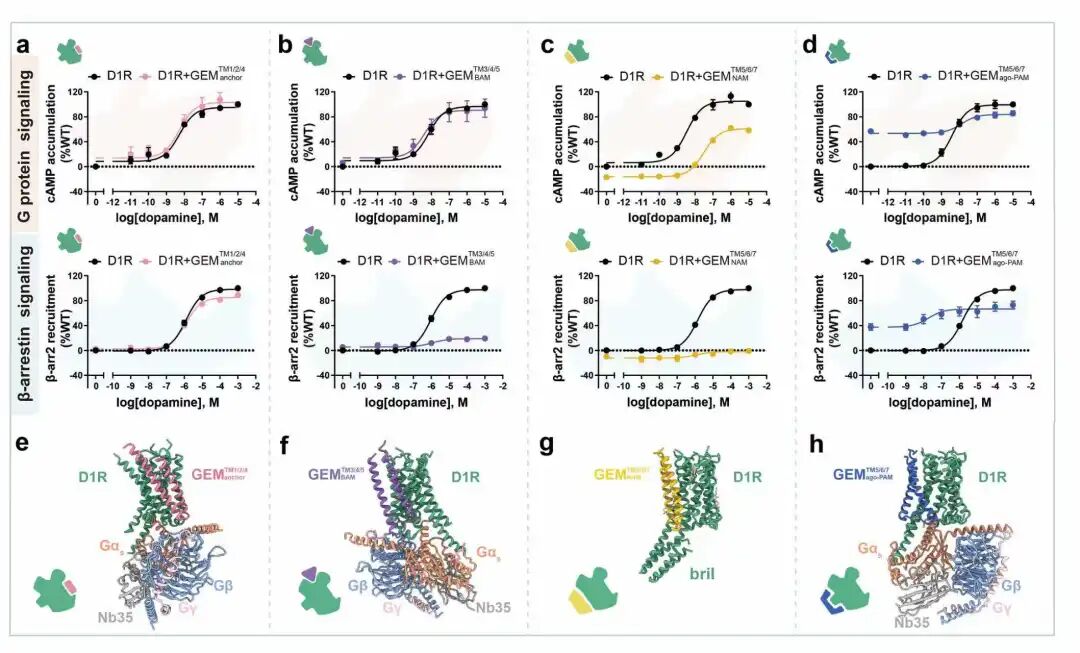
A Programmable Protein "Armor"
"At the initial stage of the research, I worked with senior lab members on structural genomics research, familiarizing myself with the properties and characteristics of GPCRs, which laid a theoretical foundation for subsequent design," says Shizhuo Cheng, a PhD candidate with a computer science background, trained through the interdisciplinary collaboration between medicine and computer science. Cheng attends three group meetings each week—at the medical lab, computer lab, and interdisciplinary lab—where he studies medical knowledge, hones his computer skills, and engages in constant brainstorming with his supervisors.
Close communication with both teams enabled him to translate the mysterious biological genetic code into logical and regular computer language. Surprisingly, the team discovered that the "controllability" this system confers to protein design far exceeds expectations.
"We have not only designed modules that can 'turn off', 'turn on', or generate biased signals in receptors, but we can even endow them with a certain degree of programmability," Min Zhang states. Based on this platform, these modules can be assembled into biological "logic gates" like designing electronic circuits in the future, enabling precise reprogramming of cellular signals.
Experimental data shows that the precision of AI design reaches an astonishing 1.0 angstrom, meaning the artificially designed protein achieves an atom-level, seamless fit with the experimentally resolved structure. The designed proteins can stabilize receptors in specific functional conformations, thereby realizing precise modulation of signal pathways.
"We named this component the exoskeleton protein—GEM," Yan Zhang says. "If GPCRs are likened to Sun Wukong, with 800 different transformations, then GEM is like the 'golden hoop'. It is not part of the receptor itself, yet it fits perfectly at the most critical position, able to restrict or guide its mode of action."
GEM rescues GPCR loss-of-function mutants
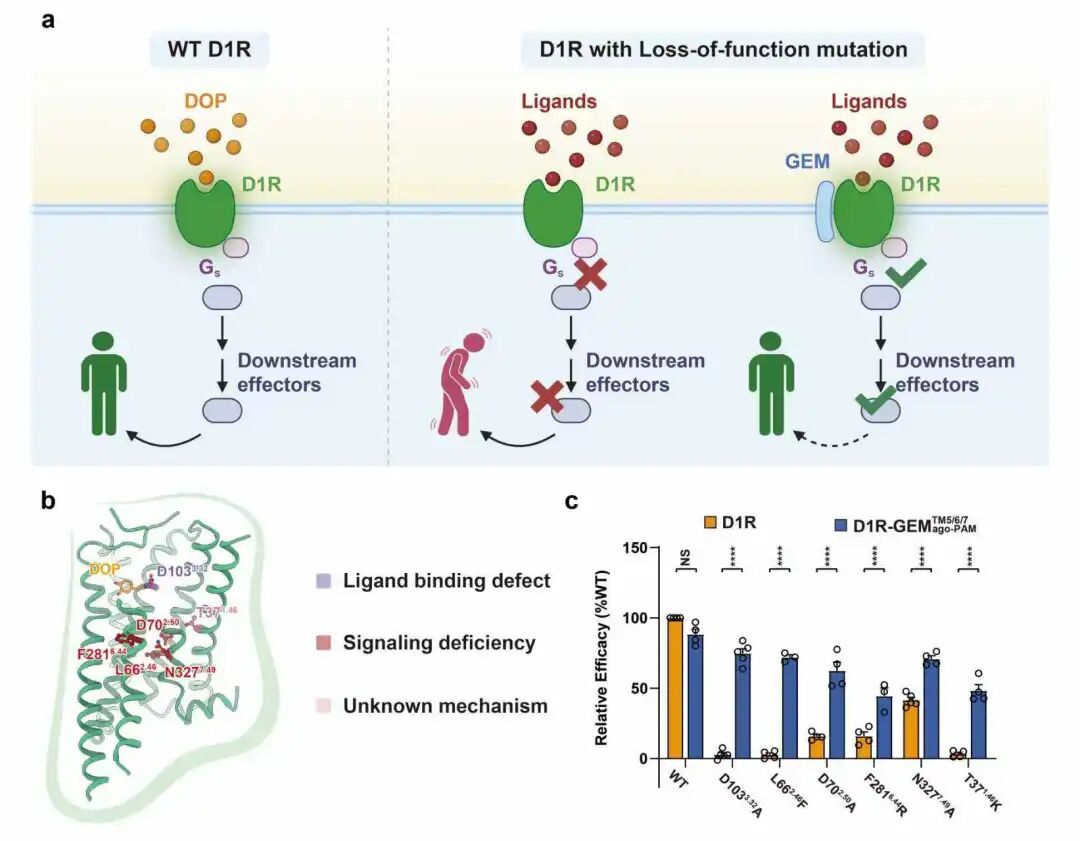
"When a receptor 'goes astray' due to mutation, becoming dysfunctional or even 'out of control', this 'golden hoop' can help it return to its normal working state; for receptors with inherently excessive activity, GEM can also exert a moderate inhibitory effect," Yan Zhang explains. "We hope this approach will provide a new therapeutic strategy for a class of GPCR dysfunction diseases that have long lacked effective intervention methods, such as certain Parkinsonism-like syndromes."
In this study, GEM represents a radical departure from traditional drug development: it provides receptors with a designable "behavioral constraint", allowing them to respond in new ways while retaining their original sensory capabilities. Simultaneously, the research establishes an AI-driven membrane protein functional design platform, offering an entirely new perspective for understanding and harnessing the body's life signal system.






